|
|
The CAPRI protein-protein docking experiment is a proven catalyst for the development of docking algorithms. An essential step in docking is the scoring of predicted binding modes generated for a given target (the experimentally determined structure to be predicted), in order to identify near-native complexes. Since 2005, the CAPRI experiment has been providing enriched data sets, including both correct and incorrect docking solutions (decoys), to enable developers to test new scoring functions independently from docking calculations.
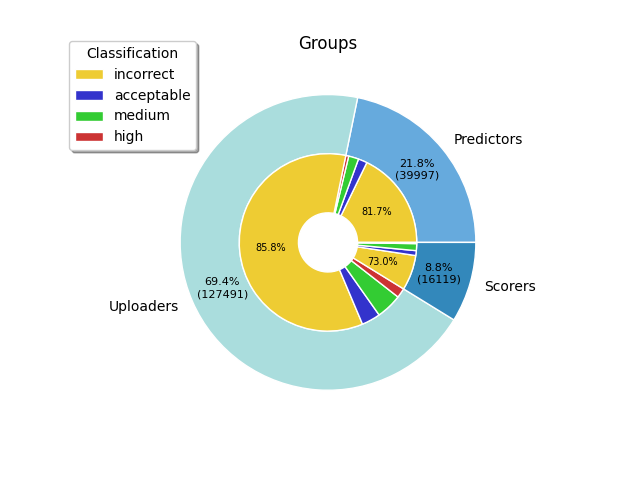
Here we present the ensemble of models submitted to the CAPRI docking and scoring experiments for CAPRI targets with published PDB structures. All models have been annotated with calculated assessment quantities used by CAPRI.
| 96 |
Targets |
| 148 |
Interfaces |
| 170310 |
Decoys |
| 57.45G |
Size |
|
|
|
|
|
|
|




